Advanced predictions, data and plotting¶
NBNode can leverage its .data attribute for prediction and plotting.
Predict based on node names¶
Sometimes it might be usefull to identify the node corresponding to its name. Usually, try to set your name such that it matches NBNode.get_name_full(), but it might be of convenience.
[1]:
import nbnode.nbnode_trees as nbtree
mytree = nbtree.tree_simple()
mytree.pretty_print()
print(mytree.prediction_str("/a"))
print(mytree.prediction_str("/a/a0"))
print(mytree.prediction_str("/a/a1"))
# In this case, probably better:
print(mytree["/a"])
print(mytree["/a/a0"])
print(mytree["/a/a1"])
# It becomes interesting when the splits are different or the root node is missing:
print(mytree.prediction_str("/a/a1/a1a"))
print(mytree["/a/a1/a1a"])
print("\nMissing root")
print(mytree.prediction_str("a/a1/a1a"))
print(mytree["a/a1/a1a"])
print("\nDifferent split")
print(mytree.prediction_str(":a:a1:a1a", split=":"))
print(mytree[":a:a1:a1a"])
a (counter:0)
├── a0 (counter:0)
├── a1 (counter:0)
│ └── a1a (counter:0)
└── a2 (counter:0)
NBNode('/a', counter=0, decision_name=None, decision_value=None)
NBNode('/a/a0', counter=0, decision_name='m1', decision_value=-1)
NBNode('/a/a1', counter=0, decision_name='m1', decision_value=1)
NBNode('/a', counter=0, decision_name=None, decision_value=None)
NBNode('/a/a0', counter=0, decision_name='m1', decision_value=-1)
NBNode('/a/a1', counter=0, decision_name='m1', decision_value=1)
NBNode('/a/a1/a1a', counter=0, decision_name='m2', decision_value='test')
NBNode('/a/a1/a1a', counter=0, decision_name='m2', decision_value='test')
Missing root
NBNode('/a/a1/a1a', counter=0, decision_name='m2', decision_value='test')
None
Different split
NBNode('/a/a1/a1a', counter=0, decision_name='m2', decision_value='test')
None
Usage of .data¶
The .data is quite powerfull for further predictions, math and plotting.
[2]:
import os
import re
import pandas as pd
print(os.getcwd())
cellmat = pd.read_csv(
os.path.join(
os.pardir, os.pardir, "tests", "testdata", "flowcytometry", "gated_cells", "cellmat.csv"
)
)
# FS TOF (against FS INT which is "FS")
cellmat.rename(columns={"FS_TOF": "FS.0"}, inplace=True)
cellmat.columns = [re.sub("_.*", "", x) for x in cellmat.columns]
print(cellmat)
/home/gugl/clonedgit/ccc_verse/nbnode/docs/notebooks
FS FS.0 SS CD45RA CCR7 CD28 PD1 CD27 CD4 CD8 CD3
0 197657 94 186372 3.90 6.34 4.97 -1.98 7.51 5.87 3.55 5.83 \
1 180716 92 135447 6.48 6.63 5.17 3.07 7.38 5.49 2.64 5.83
2 134129 90 168268 5.92 6.53 5.39 2.60 7.57 5.70 2.54 5.74
3 239241 94 79262 5.47 6.57 4.68 3.30 7.36 5.75 2.76 6.06
4 246527 89 97635 6.12 6.26 5.22 3.05 7.40 5.70 2.66 6.29
.. ... ... ... ... ... ... ... ... ... ... ...
994 176236 90 149982 6.48 -1.11 2.85 -1.55 2.28 0.59 1.70 0.39
995 191863 99 115406 6.30 5.19 3.01 2.07 -1.58 0.62 1.02 0.73
996 217752 93 124675 6.35 4.75 0.42 1.89 2.02 0.52 1.48 0.53
997 334174 97 210458 1.90 1.36 1.22 2.52 -0.72 0.59 1.03 0.75
998 308089 103 219747 6.48 -0.42 1.23 2.64 7.07 0.57 1.82 1.72
CD57 CD45
0 2.62 6.78
1 2.39 6.76
2 1.02 6.46
3 1.14 6.59
4 2.22 6.33
.. ... ...
994 4.22 6.49
995 2.69 6.22
996 2.92 6.50
997 2.98 5.38
998 2.87 6.03
[999 rows x 13 columns]
[3]:
import nbnode.nbnode_trees as nbtree
cell_tree = nbtree.tree_complete_aligned_trunk()
cell_tree.data = cellmat
# When setting .data, we do not need to give the cellmat explicitely to .predict()
cell_tree.id_preds(cell_tree.predict())
cell_tree.count(use_ids=True)
cell_tree.pretty_print()
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
Accessing any node’s data is easy and is built on rootnode._data and node.ids such that every node.data := root._data.iloc[self.ids, :].
[4]:
print(cell_tree.data.iloc[0:3, :])
print(cell_tree.data.shape)
print(cell_tree["/AllCells/CD4+/CD8-"].data.iloc[0:3, :])
print(cell_tree["/AllCells/CD4+/CD8-"].data.shape)
FS FS.0 SS CD45RA CCR7 CD28 PD1 CD27 CD4 CD8 CD3
0 197657 94 186372 3.90 6.34 4.97 -1.98 7.51 5.87 3.55 5.83 \
1 180716 92 135447 6.48 6.63 5.17 3.07 7.38 5.49 2.64 5.83
2 134129 90 168268 5.92 6.53 5.39 2.60 7.57 5.70 2.54 5.74
CD57 CD45
0 2.62 6.78
1 2.39 6.76
2 1.02 6.46
(999, 13)
FS FS.0 SS CD45RA CCR7 CD28 PD1 CD27 CD4 CD8 CD3
17 184334 107 210764 2.88 6.38 6.11 4.13 7.91 6.00 -2.65 6.15 \
25 173701 105 235028 1.81 5.53 6.40 3.83 7.28 6.01 -0.56 5.72
45 224976 101 162302 1.50 1.36 6.21 3.49 3.56 6.17 -3.40 5.37
CD57 CD45
17 2.51 6.82
25 2.66 6.80
45 1.90 6.30
(21, 13)
Now we can use the data and apply a function to it. Using it directly returns a dictionary of every node and the applied function: Dict[NBNode, Any].
[5]:
# calculate the sum of all cells per column
def col_sum(df: pd.DataFrame):
return df.apply(sum, axis=0)
# There are 13 nodes in that tree, therefore apply returns a DICTIONARY of 13 elements
print(len(cell_tree.apply(fun=col_sum)))
print("\n\n\n")
print(cell_tree.apply(fun=col_sum))
13
{NBNode('/AllCells', counter=999, decision_name=None, decision_value=None): FS 4.463310e+08
FS.0 1.077770e+05
SS 7.552312e+08
CD45RA 2.627850e+03
CCR7 2.423360e+03
CD28 2.544160e+03
PD1 2.563000e+03
CD27 2.087390e+03
CD4 1.559720e+03
CD8 2.202570e+03
CD3 1.274720e+03
CD57 3.276380e+03
CD45 5.539190e+03
dtype: float64, NBNode('/AllCells/DN', counter=0, decision_name=['CD4', 'CD8'], decision_value=[-1, -1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64, NBNode('/AllCells/DP', counter=973, decision_name=['CD4', 'CD8'], decision_value=[1, 1]): FS 4.381676e+08
FS.0 1.050100e+05
SS 7.451190e+08
CD45RA 2.549230e+03
CCR7 2.365810e+03
CD28 2.479180e+03
PD1 2.469230e+03
CD27 2.012120e+03
CD4 1.475720e+03
CD8 2.203020e+03
CD3 1.220460e+03
CD57 3.175190e+03
CD45 5.372880e+03
dtype: float64, NBNode('/AllCells/CD4-/CD8+', counter=5, decision_name=['CD4', 'CD8'], decision_value=[-1, 1]): FS 1675272.00
FS.0 520.00
SS 1620265.00
CD45RA 21.41
CCR7 10.15
CD28 2.48
PD1 15.34
CD27 16.49
CD4 -0.60
CD8 32.84
CD3 18.20
CD57 27.75
CD45 33.62
dtype: float64, NBNode('/AllCells/CD4-/CD8+/naive', counter=5, decision_name=['CCR7', 'CD45RA'], decision_value=[1, 1]): FS 1675272.00
FS.0 520.00
SS 1620265.00
CD45RA 21.41
CCR7 10.15
CD28 2.48
PD1 15.34
CD27 16.49
CD4 -0.60
CD8 32.84
CD3 18.20
CD57 27.75
CD45 33.62
dtype: float64, NBNode('/AllCells/CD4-/CD8+/Tcm', counter=0, decision_name=['CCR7', 'CD45RA'], decision_value=[1, -1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64, NBNode('/AllCells/CD4-/CD8+/Temra', counter=0, decision_name=['CCR7', 'CD45RA'], decision_value=[-1, 1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64, NBNode('/AllCells/CD4-/CD8+/Tem', counter=0, decision_name=['CCR7', 'CD45RA'], decision_value=[-1, -1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64, NBNode('/AllCells/CD4+/CD8-', counter=21, decision_name=['CD4', 'CD8'], decision_value=[1, -1]): FS 6488119.00
FS.0 2247.00
SS 8491889.00
CD45RA 57.21
CCR7 47.40
CD28 62.50
PD1 78.43
CD27 58.78
CD4 84.60
CD8 -33.29
CD3 36.06
CD57 73.44
CD45 132.69
dtype: float64, NBNode('/AllCells/CD4+/CD8-/naive', counter=20, decision_name=['CCR7', 'CD45RA'], decision_value=[1, 1]): FS 6116150.00
FS.0 2146.00
SS 8234532.00
CD45RA 51.22
CCR7 47.44
CD28 60.27
PD1 79.05
CD27 57.58
CD4 81.01
CD8 -32.36
CD3 34.95
CD57 69.94
CD45 126.00
dtype: float64, NBNode('/AllCells/CD4+/CD8-/Tcm', counter=0, decision_name=['CCR7', 'CD45RA'], decision_value=[1, -1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64, NBNode('/AllCells/CD4+/CD8-/Temra', counter=1, decision_name=['CCR7', 'CD45RA'], decision_value=[-1, 1]): FS 371969.00
FS.0 101.00
SS 257357.00
CD45RA 5.99
CCR7 -0.04
CD28 2.23
PD1 -0.62
CD27 1.20
CD4 3.59
CD8 -0.93
CD3 1.11
CD57 3.50
CD45 6.69
dtype: float64, NBNode('/AllCells/CD4+/CD8-/Tem', counter=0, decision_name=['CCR7', 'CD45RA'], decision_value=[-1, -1]): FS 0
FS.0 0
SS 0
CD45RA 0
CCR7 0
CD28 0
PD1 0
CD27 0
CD4 0
CD8 0
CD3 0
CD57 0
CD45 0
dtype: int64}
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
But it is probably often more useful to directly set a (new) attribute of each node to the result!
[6]:
# calculate the sum of all cells per column
def col_sum(df: pd.DataFrame):
return df.apply(sum, axis=0)
print("\n\n\n")
res = cell_tree.apply(fun=col_sum, result_attribute_name="col_sum")
print(res is None)
print(cell_tree.col_sum)
True
FS 4.463310e+08
FS.0 1.077770e+05
SS 7.552312e+08
CD45RA 2.627850e+03
CCR7 2.423360e+03
CD28 2.544160e+03
PD1 2.563000e+03
CD27 2.087390e+03
CD4 1.559720e+03
CD8 2.202570e+03
CD3 1.274720e+03
CD57 3.276380e+03
CD45 5.539190e+03
dtype: float64
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
We also introduce a convenience wrapper: per_node_data_fun. It first transforms the data as datatable.Frame, then potentially restricts the data to a subset of columns and finally applies attributes (functions!) of datatable.Frame. Additionally we created a function to calculate the covariance matrix (based on datatable).
[7]:
from nbnode.nbnode_util import per_node_data_fun
print("\n\n\n")
cell_tree.apply(
lambda x: per_node_data_fun(x=x, fun_name="mean"),
result_attribute_name="mean",
)
print(cell_tree.mean)
print("\n\n")
cell_tree.apply(
lambda x: per_node_data_fun(x=x, fun_name="cov"),
result_attribute_name="cov",
)
print("\n\n")
cell_tree.cov
FS FS.0 SS CD45RA CCR7 CD28
0 446777.783784 107.884885 755987.182182 2.63048 2.425786 2.546707 \
PD1 CD27 CD4 CD8 CD3 CD57 CD45
0 2.565566 2.089479 1.561281 2.204775 1.275996 3.27966 5.544735
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
[7]:
| FS | FS.0 | SS | CD45RA | CCR7 | CD28 | PD1 | CD27 | CD4 | CD8 | CD3 | CD57 | CD45 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| FS | 2.165583e+10 | 6.147502e+05 | 3.620725e+10 | -39147.927371 | -14758.007195 | 2558.966552 | 11245.428569 | -82448.944161 | -83005.474472 | -33479.754087 | -103486.452630 | 5706.132441 | -26665.567051 |
| FS.0 | 6.147502e+05 | 1.254948e+02 | 1.563292e+06 | -0.995866 | -0.464734 | -0.241061 | 1.563517 | -3.511773 | -2.612908 | -1.555762 | -5.162145 | 0.418738 | -0.143062 |
| SS | 3.620725e+10 | 1.563292e+06 | 1.060875e+11 | -126466.386170 | -36703.726977 | 10494.293897 | 20883.824316 | -232079.856598 | -223074.687248 | -114591.527895 | -284654.480683 | -20623.042663 | -101119.493078 |
| CD45RA | -3.914793e+04 | -9.958665e-01 | -1.264664e+05 | 1.377467 | 0.424056 | -0.225620 | -0.058146 | 0.531804 | 0.254024 | 0.488336 | 0.467378 | 0.423125 | 0.483976 |
| CCR7 | -1.475801e+04 | -4.647341e-01 | -3.670373e+04 | 0.424056 | 1.235744 | 0.711549 | 0.153438 | 1.180837 | 0.571201 | 0.342640 | 0.772448 | -0.124259 | 0.239509 |
| CD28 | 2.558967e+03 | -2.410609e-01 | 1.049429e+04 | -0.225620 | 0.711549 | 1.896833 | 0.177192 | 1.152222 | 0.648468 | 0.054631 | 0.807809 | -0.530344 | 0.117106 |
| PD1 | 1.124543e+04 | 1.563517e+00 | 2.088382e+04 | -0.058146 | 0.153438 | 0.177192 | 1.161104 | 0.027818 | 0.205716 | -0.008117 | 0.141911 | 0.209764 | 0.143325 |
| CD27 | -8.244894e+04 | -3.511773e+00 | -2.320799e+05 | 0.531804 | 1.180837 | 1.152222 | 0.027818 | 3.862032 | 1.129160 | 1.014295 | 1.892278 | -0.049220 | 0.553066 |
| CD4 | -8.300547e+04 | -2.612908e+00 | -2.230747e+05 | 0.254024 | 0.571201 | 0.648468 | 0.205716 | 1.129160 | 1.703458 | -0.054002 | 1.091317 | -0.102284 | 0.512471 |
| CD8 | -3.347975e+04 | -1.555762e+00 | -1.145915e+05 | 0.488336 | 0.342640 | 0.054631 | -0.008117 | 1.014295 | -0.054002 | 2.119584 | 1.205916 | 0.385426 | 0.343696 |
| CD3 | -1.034865e+05 | -5.162145e+00 | -2.846545e+05 | 0.467378 | 0.772448 | 0.807809 | 0.141911 | 1.892278 | 1.091317 | 1.205916 | 2.316320 | 0.125443 | 0.564875 |
| CD57 | 5.706132e+03 | 4.187383e-01 | -2.062304e+04 | 0.423125 | -0.124259 | -0.530344 | 0.209764 | -0.049220 | -0.102284 | 0.385426 | 0.125443 | 0.886665 | 0.296332 |
| CD45 | -2.666557e+04 | -1.430616e-01 | -1.011195e+05 | 0.483976 | 0.239509 | 0.117106 | 0.143325 | 0.553066 | 0.512471 | 0.343696 | 0.564875 | 0.296332 | 0.605821 |
Importantly, the apply function is done on all nodes separately, therefore also the result_attribute_name is set for each node separately!
[8]:
cell_tree.pretty_print()
cell_tree["/AllCells/CD4+/CD8-"].cov
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
[8]:
| FS | FS.0 | SS | CD45RA | CCR7 | CD28 | PD1 | CD27 | CD4 | CD8 | CD3 | CD57 | CD45 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| FS | 8.579379e+09 | -70918.9500 | 1.983754e+10 | 55738.251286 | -47391.339857 | -70431.262810 | 62951.568762 | -105445.765952 | -108395.155929 | -5903.629738 | -168594.759857 | 76528.430143 | -23592.250929 |
| FS.0 | -7.091895e+04 | 161.5000 | 6.775819e+05 | 4.836000 | 0.256000 | -5.529000 | 5.402000 | -13.373000 | -2.360500 | -0.549000 | -8.939000 | 2.551000 | 0.333000 |
| SS | 1.983754e+10 | 677581.9500 | 7.257807e+10 | 78956.079500 | 7015.310000 | -61653.769333 | 381992.605667 | -65083.371833 | -297815.850500 | -87121.889333 | -440940.419500 | 283685.350500 | -83679.807500 |
| CD45RA | 5.573825e+04 | 4.8360 | 7.895608e+04 | 1.984566 | -0.231057 | -0.625068 | 0.014314 | -1.343086 | -0.634414 | -0.121121 | -1.264832 | 0.641103 | 0.006621 |
| CCR7 | -4.739134e+04 | 0.2560 | 7.015310e+03 | -0.231057 | 2.119731 | 1.837819 | 1.227909 | 2.731482 | 0.721181 | -0.526321 | 1.609736 | 0.085096 | 0.045586 |
| CD28 | -7.043126e+04 | -5.5290 | -6.165377e+04 | -0.625068 | 1.837819 | 4.233845 | 1.408394 | 4.528386 | 1.430229 | -1.169376 | 3.352144 | -0.117211 | -0.155401 |
| PD1 | 6.295157e+04 | 5.4020 | 3.819926e+05 | 0.014314 | 1.227909 | 1.408394 | 4.538896 | 1.954455 | -1.065938 | -1.859649 | -1.890051 | 2.319644 | -0.639173 |
| CD27 | -1.054458e+05 | -13.3730 | -6.508337e+04 | -1.343086 | 2.731482 | 4.528386 | 1.954455 | 8.181249 | 1.576484 | -1.738945 | 3.635942 | 0.463187 | -0.032486 |
| CD4 | -1.083952e+05 | -2.3605 | -2.978159e+05 | -0.634414 | 0.721181 | 1.430229 | -1.065938 | 1.576484 | 1.823103 | 0.092512 | 2.923721 | -1.296814 | 0.386778 |
| CD8 | -5.903630e+03 | -0.5490 | -8.712189e+04 | -0.121121 | -0.526321 | -1.169376 | -1.859649 | -1.738945 | 0.092512 | 1.651106 | 0.266684 | -1.010761 | 0.280202 |
| CD3 | -1.685948e+05 | -8.9390 | -4.409404e+05 | -1.264832 | 1.609736 | 3.352144 | -1.890051 | 3.635942 | 2.923721 | 0.266684 | 5.852411 | -2.329204 | 0.543411 |
| CD57 | 7.652843e+04 | 2.5510 | 2.836854e+05 | 0.641103 | 0.085096 | -0.117211 | 2.319644 | 0.463187 | -1.296814 | -1.010761 | -2.329204 | 1.865231 | -0.371314 |
| CD45 | -2.359225e+04 | 0.3330 | -8.367981e+04 | 0.006621 | 0.045586 | -0.155401 | -0.639173 | -0.032486 | 0.386778 | 0.280202 | 0.543411 | -0.371314 | 0.261683 |
Finally, apply() does not need to use .data but can use any attribute present on all nodes.
[9]:
import nbnode.nbnode_trees as nbtree
cell_tree = nbtree.tree_complete_aligned_trunk()
cell_tree.data = cellmat
# When setting .data, we do not need to give the cellmat explicitely to .predict()
cell_tree.id_preds(cell_tree.predict())
cell_tree.count(use_ids=True)
for x in cell_tree.__dict__.keys():
print(x)
print("\n\n")
name
_NodeMixin__children
decision_name
decision_value
decision_cutoff
counter
ids
_data
id_unique_dot_exporter
math_node_attribute
math_inplace
_long_print_attributes
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
warnings.warn(
[10]:
try:
print(cell_tree.changed_decision_name)
except AttributeError as e:
print("AttributeError: 'NBNode' object has no attribute 'changed_decision_name'")
cell_tree.apply(lambda x: x[0] + "___this_is_a_change" if x is not None else x, input_attribute_name="decision_name", result_attribute_name="changed_decision_name")
cell_tree.pretty_print(print_attributes=["decision_name", "changed_decision_name"])
AttributeError: 'NBNode' object has no attribute 'changed_decision_name'
AllCells (decision_name:None, changed_decision_name:None)
├── DN (decision_name:['CD4', 'CD8'], changed_decision_name:CD4___this_is_a_change)
├── DP (decision_name:['CD4', 'CD8'], changed_decision_name:CD4___this_is_a_change)
├── CD4-/CD8+ (decision_name:['CD4', 'CD8'], changed_decision_name:CD4___this_is_a_change)
│ ├── naive (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
│ ├── Tcm (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
│ ├── Temra (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
│ └── Tem (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
└── CD4+/CD8- (decision_name:['CD4', 'CD8'], changed_decision_name:CD4___this_is_a_change)
├── naive (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
├── Tcm (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
├── Temra (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
└── Tem (decision_name:['CCR7', 'CD45RA'], changed_decision_name:CCR7___this_is_a_change)
Plotting¶
For plotting we have two major steps:
Generate a
`.dot<https://graphviz.org/doc/info/lang.html>`__ file with anytreeUse
`graphviz<https://graphviz.org/download/>`__ as engine to make the plots
Potentially you have to install graphviz.
[11]:
import nbnode.nbnode_trees as nbtree
cell_tree = nbtree.tree_complete_aligned_trunk()
exported_dot = cell_tree.export_dot()
print(exported_dot)
digraph tree {
node [shape=box, style="filled", color="black"];
"0x7f6217352b80" [label="None", fillcolor="white"];
"0x7f628c31cfa0" [label="None", fillcolor="white"];
"0x7f628c31c9d0" [label="None", fillcolor="white"];
"0x7f628c31cb20" [label="None", fillcolor="white"];
"0x7f6217352670" [label="None", fillcolor="white"];
"0x7f6217352730" [label="None", fillcolor="white"];
"0x7f6217352610" [label="None", fillcolor="white"];
"0x7f6217352820" [label="None", fillcolor="white"];
"0x7f6217352d00" [label="None", fillcolor="white"];
"0x7f6217352190" [label="None", fillcolor="white"];
"0x7f6217352850" [label="None", fillcolor="white"];
"0x7f62173528b0" [label="None", fillcolor="white"];
"0x7f62173528e0" [label="None", fillcolor="white"];
"0x7f6217352b80" -> "0x7f628c31cfa0";
"0x7f6217352b80" -> "0x7f628c31c9d0";
"0x7f6217352b80" -> "0x7f628c31cb20";
"0x7f6217352b80" -> "0x7f6217352d00";
"0x7f628c31cb20" -> "0x7f6217352670";
"0x7f628c31cb20" -> "0x7f6217352730";
"0x7f628c31cb20" -> "0x7f6217352610";
"0x7f628c31cb20" -> "0x7f6217352820";
"0x7f6217352d00" -> "0x7f6217352190";
"0x7f6217352d00" -> "0x7f6217352850";
"0x7f6217352d00" -> "0x7f62173528b0";
"0x7f6217352d00" -> "0x7f62173528e0";
}
[12]:
print(cell_tree.graph_from_dot())
from nbnode.plot.utils import plot_save_unified
plot_save_unified(cell_tree.graph_from_dot(), "cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
<pydotplus.graphviz.Dot object at 0x7f62141db820>
[12]:
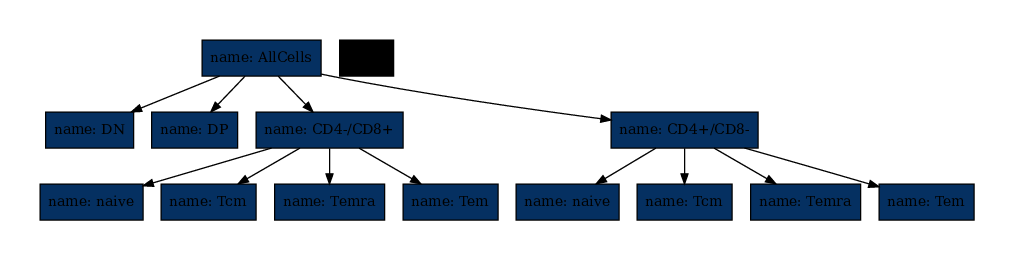
Classically, the colors visualize the number of cells in each node (counter). Up to this point we did not count, so let’s do that.
[13]:
cell_tree.data = cellmat
# When setting .data, we do not need to give the cellmat explicitely to .predict()
cell_tree.id_preds(cell_tree.predict())
cell_tree.count(use_ids=True)
cell_tree.pretty_print()
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
/home/gugl/.conda_envs/nbnode_pyscaffold/lib/python3.8/site-packages/nbnode/nbnode.py:353: UserWarning: self.ids was an empty list, subset an empty dataframe. Did you call celltree.id_preds(predicted_nodes)? Can also be a node with no cells.
[14]:
plot_save_unified(cell_tree.graph_from_dot(), "cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
[14]:
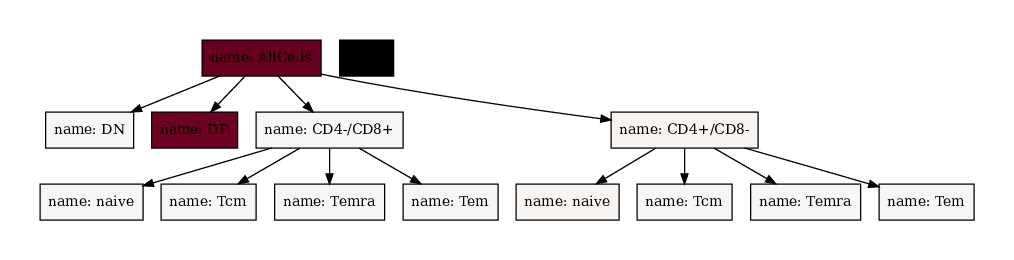
All cells are in the root node (/AllCells), therefore it is deep red. Also there are so many DP cells that all other nodes are essentially white. Let’s force maximum saturation (red) at value 50 and minimum saturation (white) at value 0!
[15]:
cell_tree.pretty_print()
plot_save_unified(
cell_tree.graph_from_dot(
fillcolor_node_attribute="counter",
custom_min_max_dict={"min": 0, "max": 50},
),
"cell_tree.pdf",
)
from wand.image import Image as WImage
img = WImage(filename="cell_tree.pdf")
img
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
[15]:
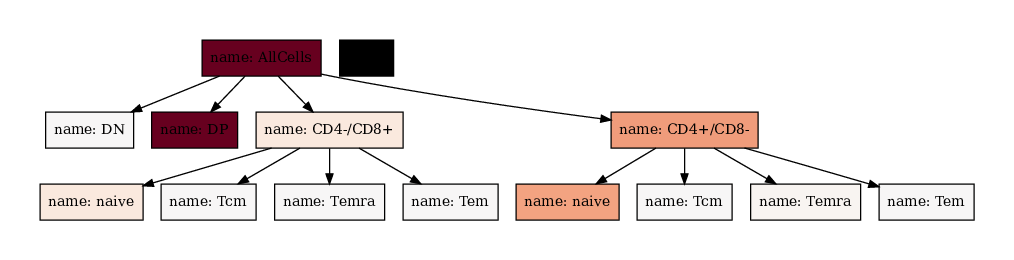
We can also modify the text printed with the node_text_attributes argument. If only a list of elements is given, the result is always value_name:value. To make more sophisticated formatting you can supply a Dict[str, str] which contains the attribute name as key and the `f-string <https://note.nkmk.me/en/python-f-strings/>`__ as value.
[16]:
cell_tree.pretty_print()
plot_save_unified(
cell_tree.graph_from_dot(
fillcolor_node_attribute="counter",
custom_min_max_dict={"min": 0, "max": 50},
node_text_attributes=["name", "counter"],
),
"cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
[16]:
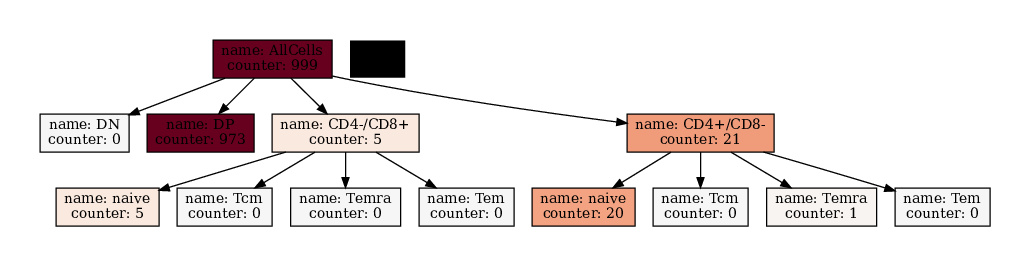
[17]:
cell_tree.pretty_print()
plot_save_unified(
cell_tree.graph_from_dot(
fillcolor_node_attribute="counter",
custom_min_max_dict={"min": 0, "max": 50},
node_text_attributes={"name":"new_format_left\n{:<8}", "counter":"new_zerofilled\n{:04d}"},
),
"cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
AllCells (counter:999)
├── DN (counter:0)
├── DP (counter:973)
├── CD4-/CD8+ (counter:5)
│ ├── naive (counter:5)
│ ├── Tcm (counter:0)
│ ├── Temra (counter:0)
│ └── Tem (counter:0)
└── CD4+/CD8- (counter:21)
├── naive (counter:20)
├── Tcm (counter:0)
├── Temra (counter:1)
└── Tem (counter:0)
[17]:
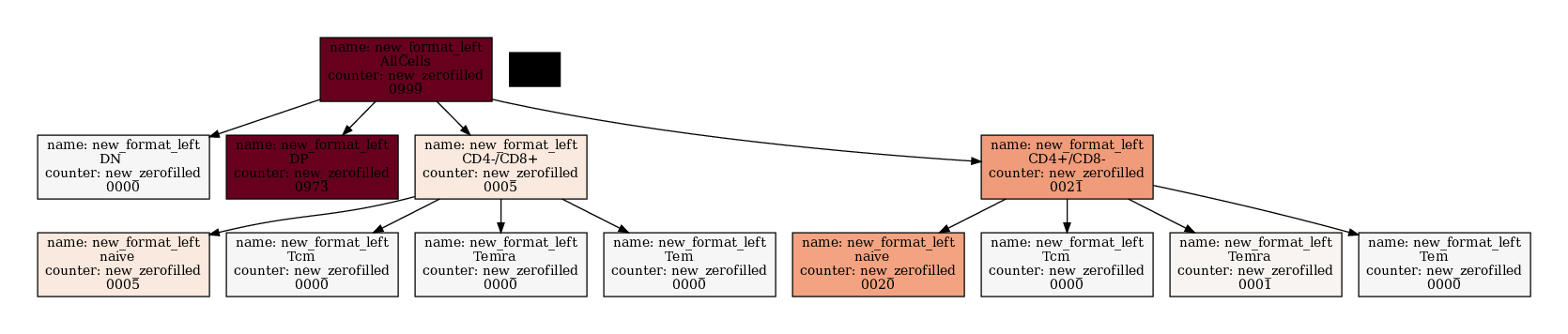
If you want to apply math to the nodes you have to do that outside.
[18]:
new_tree = (cell_tree + 1)
new_tree.astype_math_node_attribute(float)
new_tree = new_tree * 5.713223
new_tree.pretty_print()
plot_save_unified(
new_tree.graph_from_dot(
fillcolor_node_attribute="counter",
custom_min_max_dict={"min": 0, "max": 50},
node_text_attributes={"name":"new_format_left\n{:<8}", "counter":"new_zerofilled\n{:.3f}"},
),
"cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
AllCells (counter:5713.223)
├── DN (counter:5.713223)
├── DP (counter:5564.679202)
├── CD4-/CD8+ (counter:34.279338)
│ ├── naive (counter:34.279338)
│ ├── Tcm (counter:5.713223)
│ ├── Temra (counter:5.713223)
│ └── Tem (counter:5.713223)
└── CD4+/CD8- (counter:125.690906)
├── naive (counter:119.977683)
├── Tcm (counter:5.713223)
├── Temra (counter:11.426446)
└── Tem (counter:5.713223)
[18]:
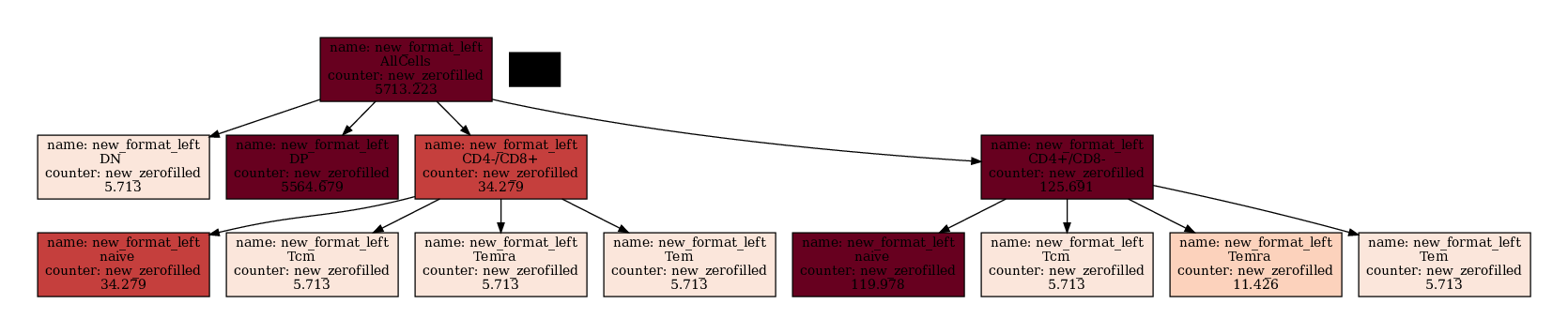
You can also change the color map also using matplotlibs color maps.
[20]:
import matplotlib as mpl
plot_save_unified(
new_tree.graph_from_dot(
fillcolor_node_attribute="counter",
cmap=mpl.colormaps.get_cmap("viridis"),
),
"cell_tree.pdf",
)
from wand.image import Image as WImage
img = WImage(filename="cell_tree.pdf")
img
[20]:
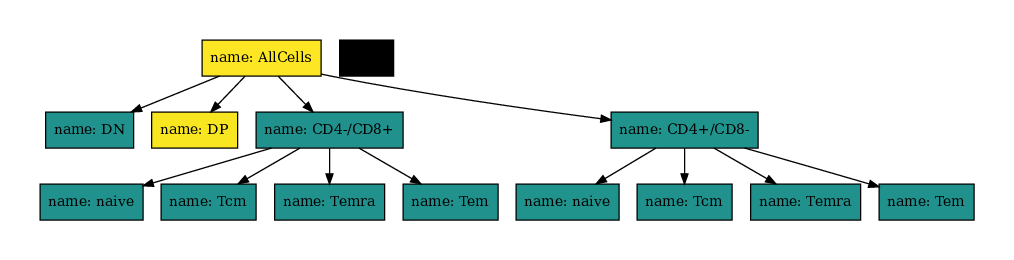
You can use usual pydotplus to make much more complex graphs:
[58]:
import pydotplus
import anytree.exporter as a_exp
from nbnode.plot.utils import plot_save_unified
simpletree = nbtree.tree_simple()
dot_data = a_exp.UniqueDotExporter(
simpletree,
options=['node [shape=box, style="filled", color="black"];'],
nodeattrfunc=lambda node: 'label="{}", fillcolor="white"'.format(node.name),
)
dotdata_str = "\n".join([x for x in dot_data])
# print(dotdata_str)
graph: pydotplus.Dot = pydotplus.graph_from_dot_data(dotdata_str)
nodes = graph.get_node_list()
plot_save_unified(
any_plot=graph, file="cell_tree.pdf"
)
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
[58]:
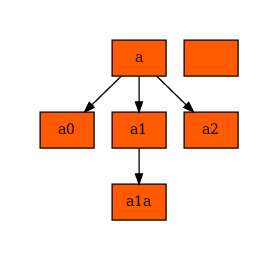
[59]:
for node in nodes:
if node.get_name() not in ("node", "edge", "plottitle"):
node.set_fillcolor("#ff5a00")
# print(graph.to_string())
plot_save_unified(any_plot=graph, file="cell_tree.pdf")
from wand.image import Image as WImage
img = WImage(filename='cell_tree.pdf')
img
[59]: