library(restrictedROC)
size_factor <- .33
main_plotname <- "res/paper/roc"
dir.create("res/paper", recursive = TRUE)
#> Warning in dir.create("res/paper", recursive = TRUE): 'res/paper' already
#> exists
# 1. Random
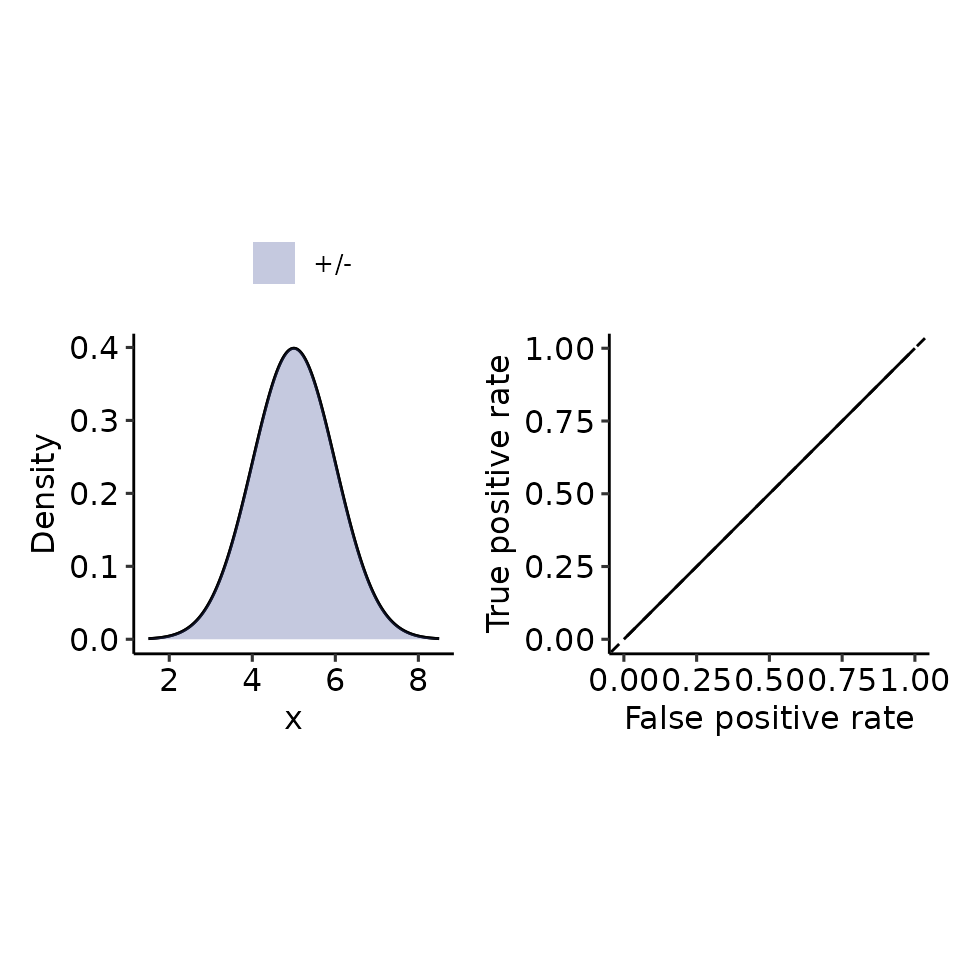
tmp <- plot_density_ROC_str(
dist_positive_str = "norm",
dist_negative = "norm",
dist_positive_args = list("mean" = 5, "sd" = 1),
dist_negative_args = list("mean" = 5, "sd" = 1),
xmin = 1.5, xmax = 8.5
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.5 with absolute error < 5.6e-15
# pdf(file.path(paste0(main_plotname, "_random.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.5 with absolute error < 5.6e-15
# dev.off()
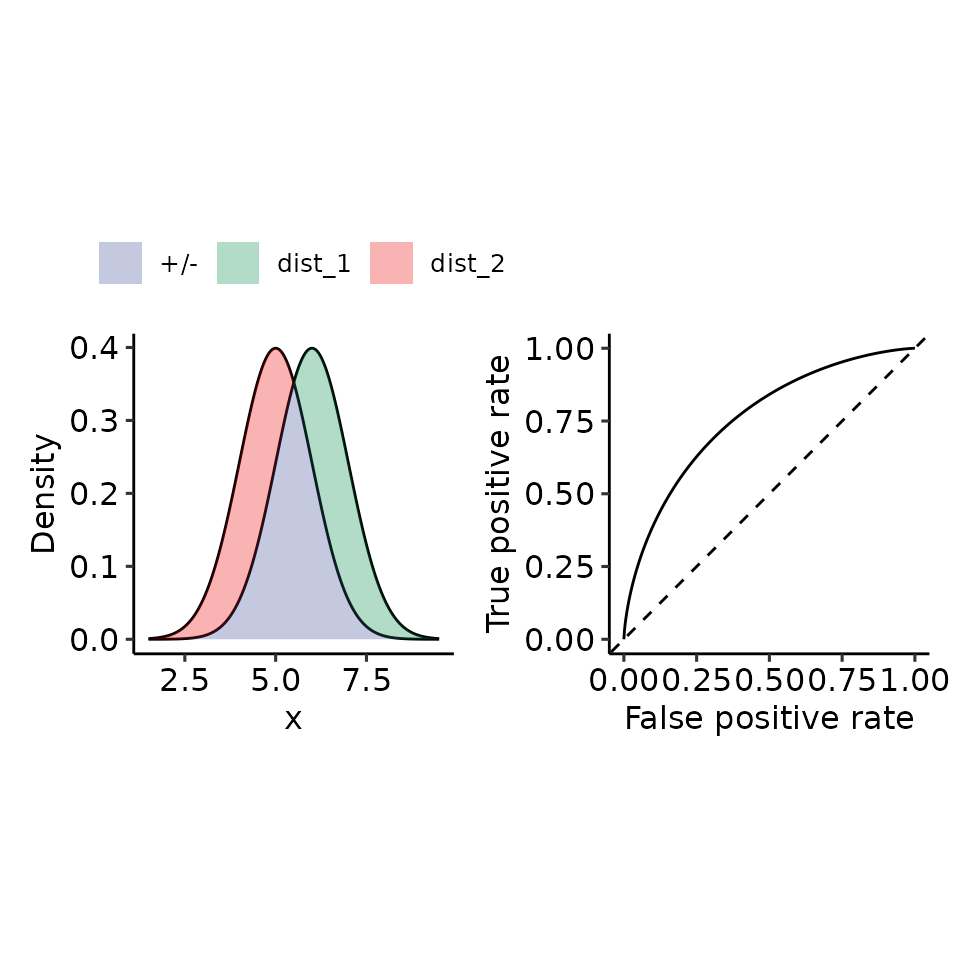
tmp <- plot_density_ROC_str(
dist_positive_str = "norm",
dist_negative = "norm",
dist_positive_args = list("mean" = 6, "sd" = 1),
dist_negative_args = list("mean" = 5, "sd" = 1),
xmin = 1.5, xmax = 9.5
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.76025 with absolute error < 4.9e-05
# pdf(file.path(paste0(main_plotname, "_posGTneg.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.76025 with absolute error < 4.9e-05
# dev.off()
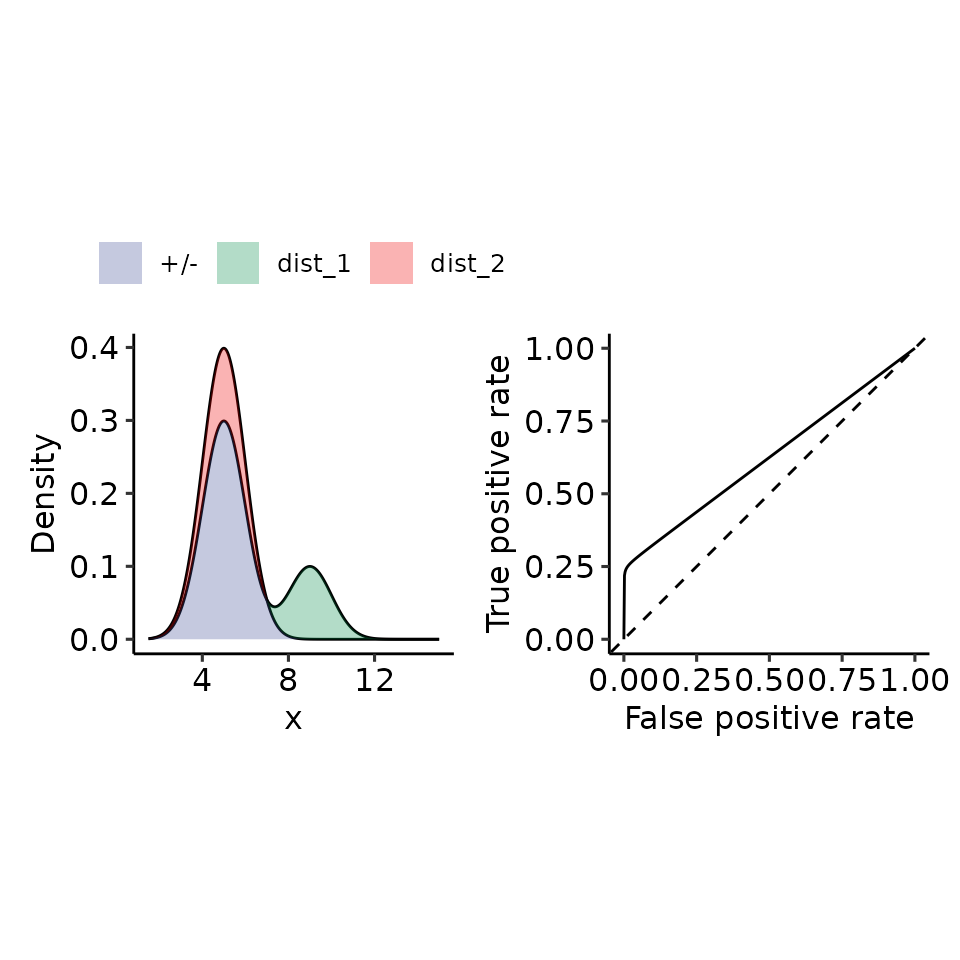
tmp <- plot_density_ROC(
density_positive = function(x) {
# .5 because the integral over each norm = 1, therefore for 2 densities I must half the resulting density.
.75 * dnorm(x, mean = 5, sd = 1) + .25 * dnorm(x, mean = 9, sd = 1)
},
cdf_positive = function(x) {
.75 * pnorm(x, mean = 5, sd = 1) + .25 * pnorm(x, mean = 9, sd = 1)
},
density_negative = function(x) dnorm(x, mean = 5, sd = 1),
quantile_negative = function(x) qnorm(x, mean = 5, sd = 1),
xmin = 1.5, xmax = 15
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.6244153 with absolute error < 1e-06
# pdf(file.path(paste0(main_plotname, "_pos2norm_highdiff.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.6244153 with absolute error < 1e-06
# dev.off()
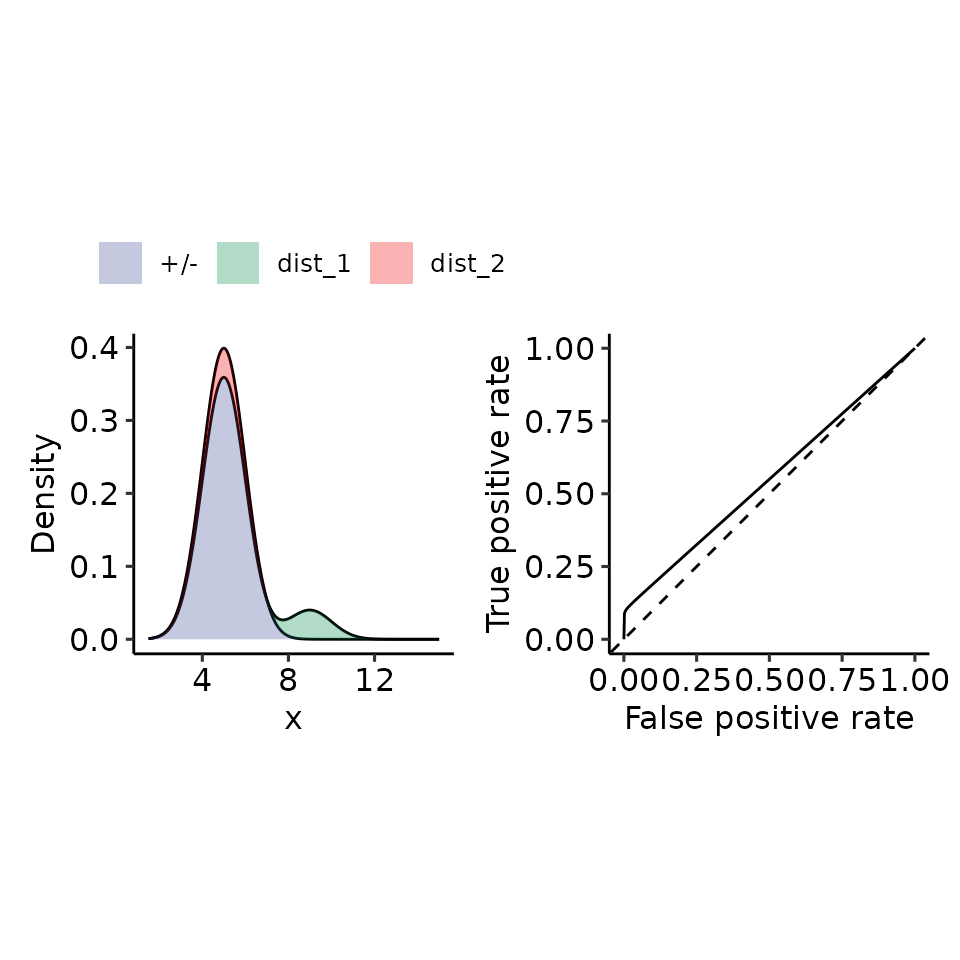
tmp <- plot_density_ROC(
density_positive = function(x) {
# .5 because the integral over each norm = 1, therefore for 2 densities I must half the resulting density.
.9 * dnorm(x, mean = 5, sd = 1) + .1 * dnorm(x, mean = 9, sd = 1)
},
cdf_positive = function(x) {
.9 * pnorm(x, mean = 5, sd = 1) + .1 * pnorm(x, mean = 9, sd = 1)
},
density_negative = function(x) dnorm(x, mean = 5, sd = 1),
quantile_negative = function(x) qnorm(x, mean = 5, sd = 1),
xmin = 1.5, xmax = 15
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.5497661 with absolute error < 4.1e-07
# pdf(file.path(paste0(main_plotname, "_pos2norm_highdiff_v2.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.5497661 with absolute error < 4.1e-07
# dev.off()
# 4. Different mean + variance
# 4.1 mean: positive > negative, var: positive > negative --> left-skewed
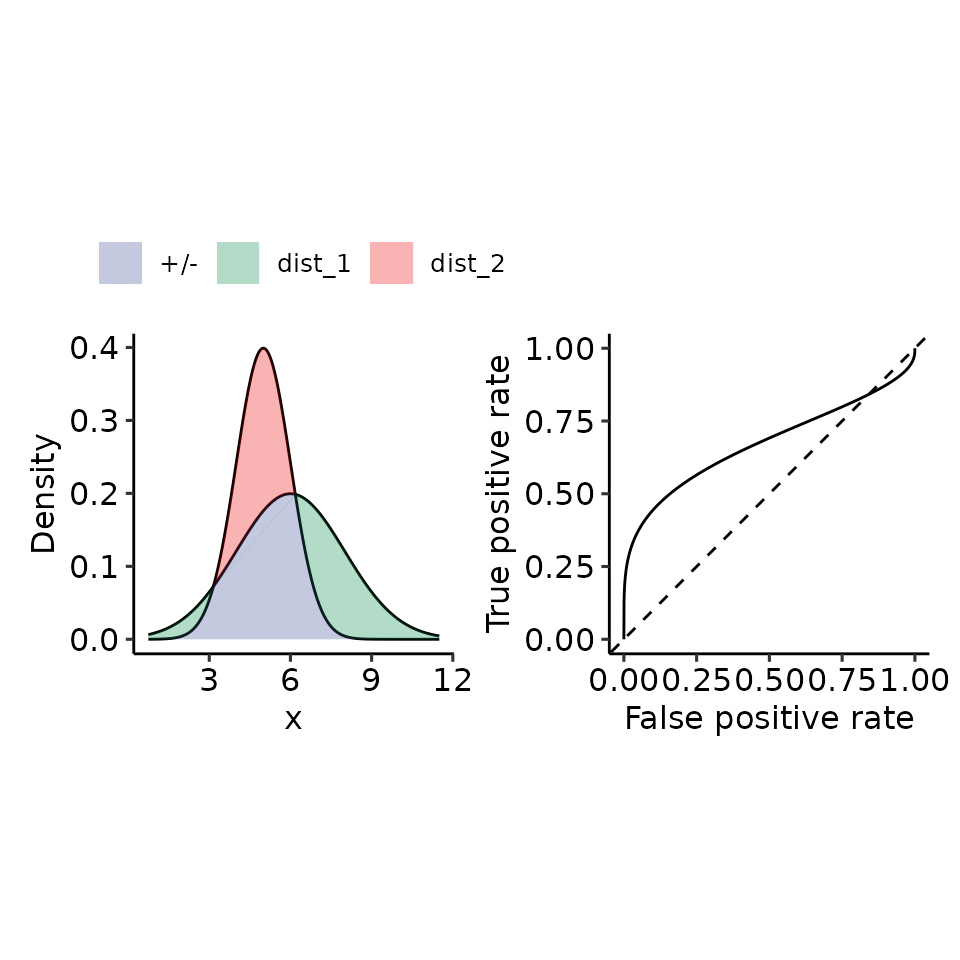
tmp <- plot_density_ROC_str(
dist_positive_str = "norm",
dist_negative = "norm",
dist_positive_args = list("mean" = 6, "sd" = 2),
dist_negative_args = list("mean" = 5, "sd" = 1),
xmin = 0.75, xmax = 11.5, length.out = 2001
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.6726396 with absolute error < 1.3e-08
# pdf(file.path(paste0(main_plotname, "_posGTneg_posVARGTneg.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.6726396 with absolute error < 1.3e-08
# dev.off()
# 4.2 mean: positive > negative, var: positive < negative --> right-skewed
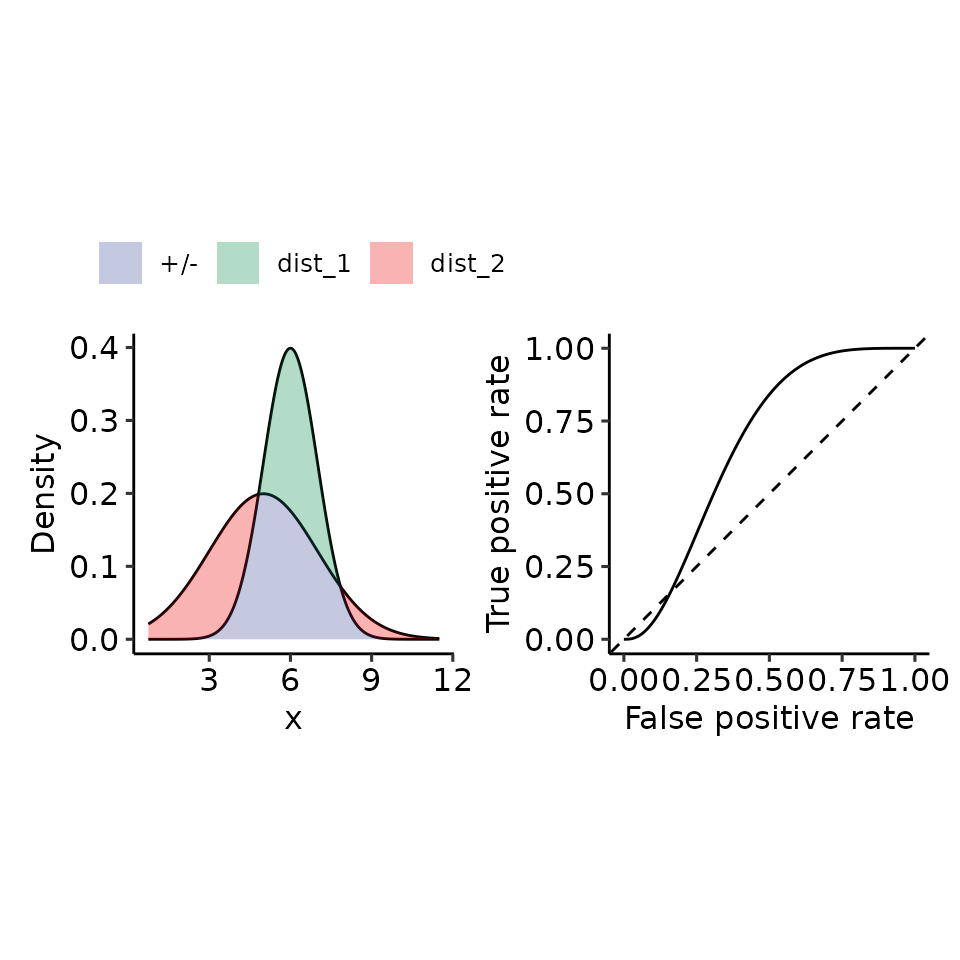
tmp <- plot_density_ROC_str(
dist_positive_str = "norm",
dist_negative = "norm",
dist_positive_args = list("mean" = 6, "sd" = 1),
dist_negative_args = list("mean" = 5, "sd" = 2),
xmin = 0.75, xmax = 11.5, length.out = 2001
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 0.6726396 with absolute error < 8.8e-07
# pdf(file.path(paste0(main_plotname, "_posGTneg_posVARLTneg.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 0.6726396 with absolute error < 8.8e-07
# dev.off()
# 5. Perfect
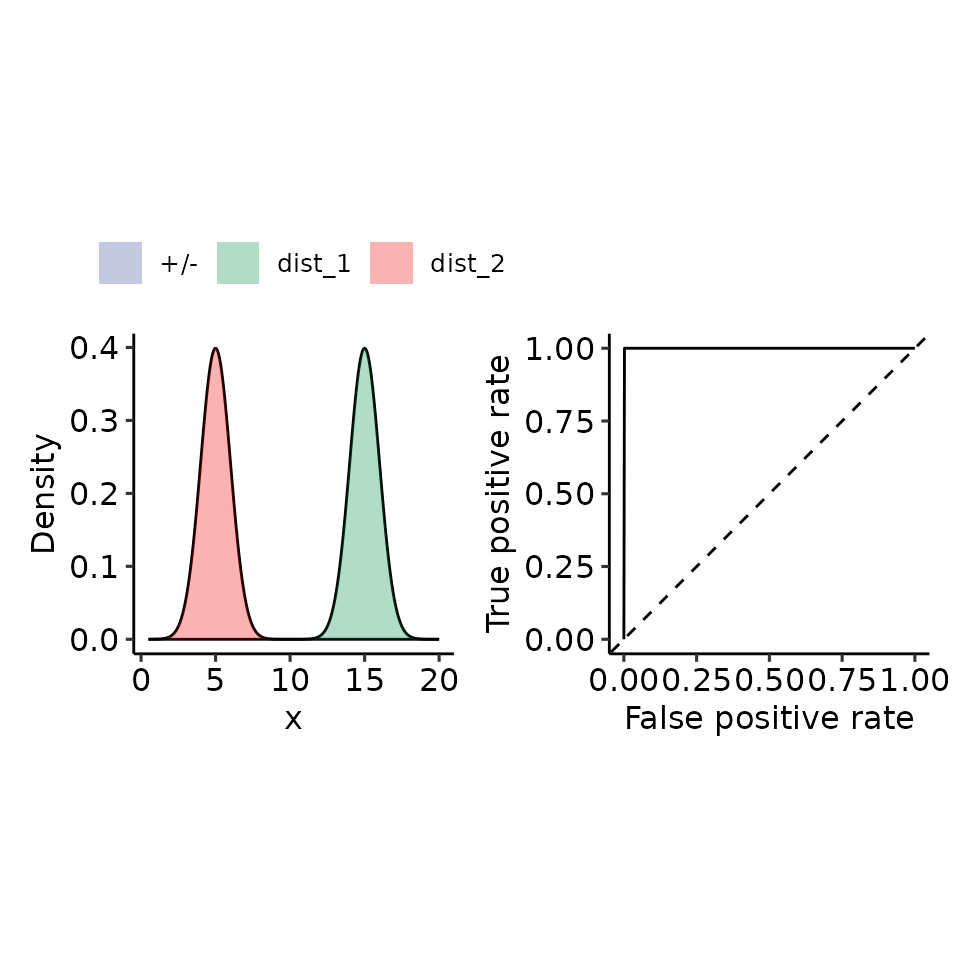
tmp <- plot_density_ROC_str(
dist_positive_str = "norm",
dist_negative = "norm",
dist_positive_args = list("mean" = 15, "sd" = 1),
dist_negative_args = list("mean" = 5, "sd" = 1),
xmin = .5, xmax = 20
)
print(tmp)
#> $plot
#> Ignoring unknown labels:
#> • colour : ""
#>
#> $auc
#> 1 with absolute error < 1.1e-14
# pdf(file.path(paste0(main_plotname, "_perfect.pdf")), height = 10 * size_factor, width = 20 * size_factor)
# print(tmp)
# # $auc
# # 1 with absolute error < 1.1e-14
# dev.off()






